
Applications are being accepted and reviewed on a rolling basis until April 15.
In order to allow time for visa processing, international scholars in particular should submit their applications as early as possible.
Course Subject:
Animals interact with the world through movement. Crawling, swimming, walking, and flying animals face different challenges. Using a range of experimental techniques, we will explore locomotor behavior, considering contributions from the nervous system and musculature, and confronting biomechanical constraints for creatures with 0 to 8 limbs in environmental conditions ranging from sea to sky.
Experiments will be organized in team-taught modules based on comparative analysis of between tractable model organisms with different means of locomotion.
Course Directors:
Instructors:
Serena Ding (MPI, Konstanz), Pavan Ramdya (EPFL), Jane Loveless (OIST), Jimmy Liao (Whitney Lab, U. Florida), Jessica Fox (Case Western Reserve), Eugenia Chiappe (CNP), Ellie Heckscher (U. Chicago), Tanya Tabachnik (Zuckerman Inst., Columbia), Andrew Straw (U. Freiburg), and Brad Dickerson (Princeton)
Guest Lecturers (preliminary list):
The summer course is closely linked to the concurrent KITP program Neurophysics of Locomotion. Course participants will attend the program's daily research seminars as part of the course curriculum. Students and lecturers will also have frequent opportunities for less formal interactions. Confirmed program participants include Jimena Berni (U. Sussex), Barry Dickson (U. Queensland), Akinao Nose (U. Tokyo), Dan Tracey (U. Indiana), Dmitri Chklovskii (Flatiron Inst. & NYU), Andrew Leifer (Princeton), David Schoppik (NYU), John Tuthill (U. Washington), Michael Dickinson (Caltech), Kathy Nagel (NYU), Simon Sponberg (Georgia Tech), Madhu Venkadesan (Yale), Ilya Nemenman (Emory), Greg Stephens (VU Amsterdam & OIST).
Course Structure
The structure for the five-week school follows the model established in the 2018 course on sensory navigation. The course will be closely linked to the concurrent Neurophysics of Locomotion program, with students attending program talks each morning, as well as joint social events. Interaction with program speakers will also be facilitated through tutorial sessions and informal meetings over coffee and meals. Students will be matched to experimental modules based on their ranked choices and have ample room to explore their own interests, based on initial instructor-guided projects. Students in different modules will interact informally – many research areas benefit from comparison or use similar technical approaches. Students teach each other, sharing their different expertise--project groups will intentionally include students from different academic backgrounds--and work closely with expert instructors, TAs. and guest lecturers from the associated KITP program.
There is no course fee, and students are provided with housing at UCSB's Santa Ynez apartments and a 14-meal plan at UCSB dining halls. A typical day's schedule includes:
- 2 morning talks at KITP
- Group lunch in a dining hall, or pizza at KITP with informal science discussion or tutorials on topics of interest
- Afternoon lab work
- Dinner in a dining hall, followed by evening lab work and science discussions
On weekends, students can attend program BBQs and self-organize outings to the beach or around Santa Barbara.
Experimental Projects (partial list)
Monika Scholz and Serena Ding: Caenorhabditis elegans locomotion in environments with different physical properties
Locomotion is intimately connected to the substrate on or in which it occurs. A swimmer in a liquid environment with drag needs to perform different strokes to effectively move its body than a walker on a hard surface. Using the roundworm C. elegans as a model, we will explore the interaction between the nervous system, muscular contractions and different substrates of locomotion in 2D and 3D motion. By altering the physical environment that the animals move in, we will explore how locomotion patterns and the underlying neural signals are altered by the environment using behavioral tracking and calcium imaging.
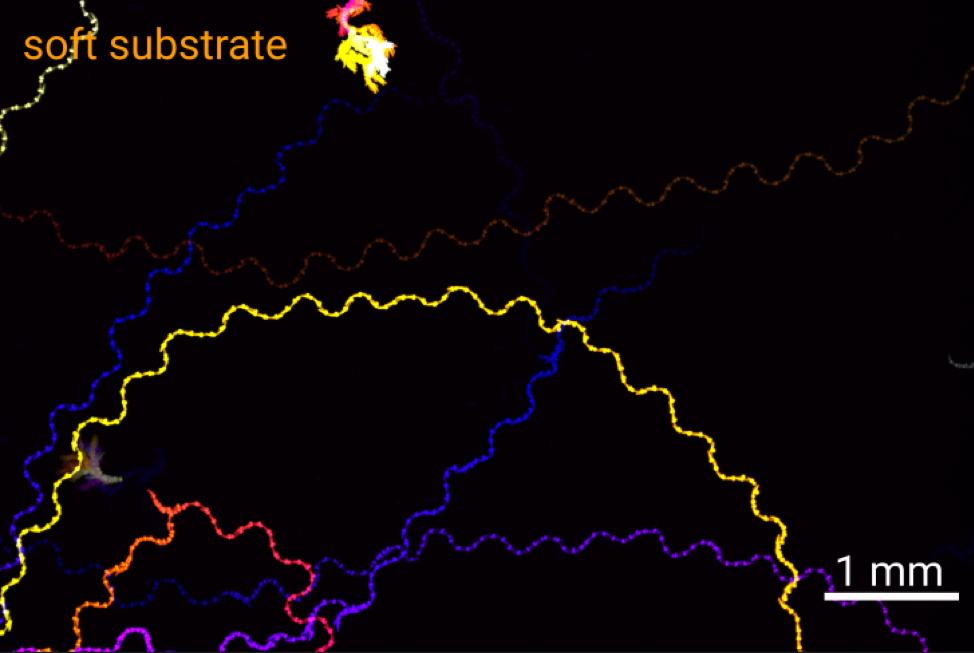

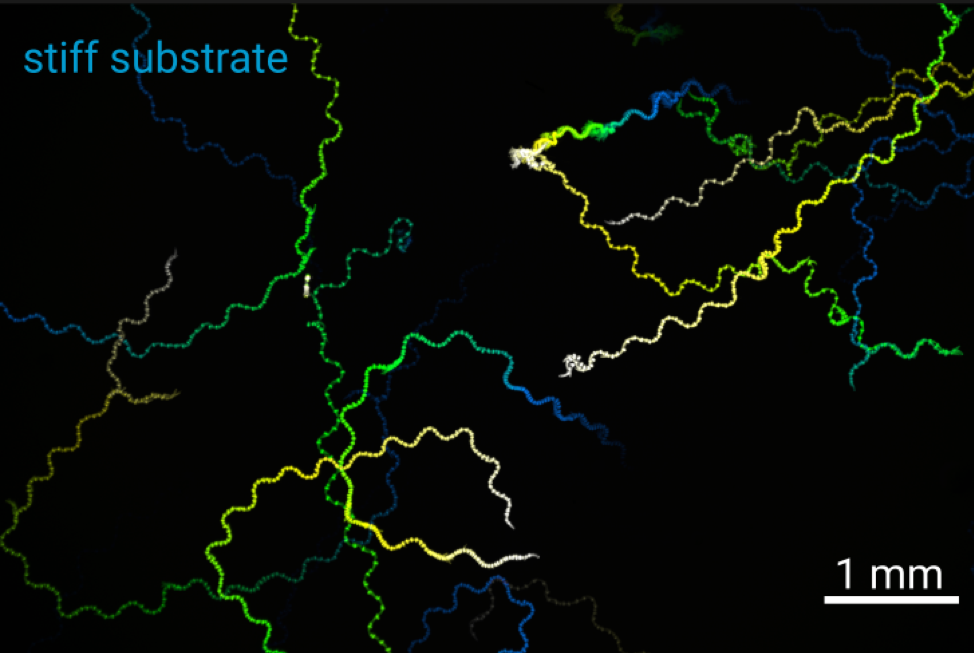
Figure: Tracks of C. elegans on soft (2% agarose) and stiff (10% agarose) substrates.
___________________________________________________________________________________________________________
Jessica Fox and Brad Dickerson: Sensory integration for fly flight
Locomotion requires animals to integrate external signals from the environment with internal signals about their own head and body movements. This is a challenge for walking animals, and becomes even more complex during high-speed, three-dimensional flight. We will use quantitative behavioral analysis, optogenetic manipulations, and neural imaging to study how fruit flies (Drosophila melanogaster) integrate visual and mechanosensory signals to control their flight. Because both vision and mechanosensation are active senses during fly flight, we will explore neuromuscular control in both the head (for gaze control) and the haltere (for mechanosensory feedback).
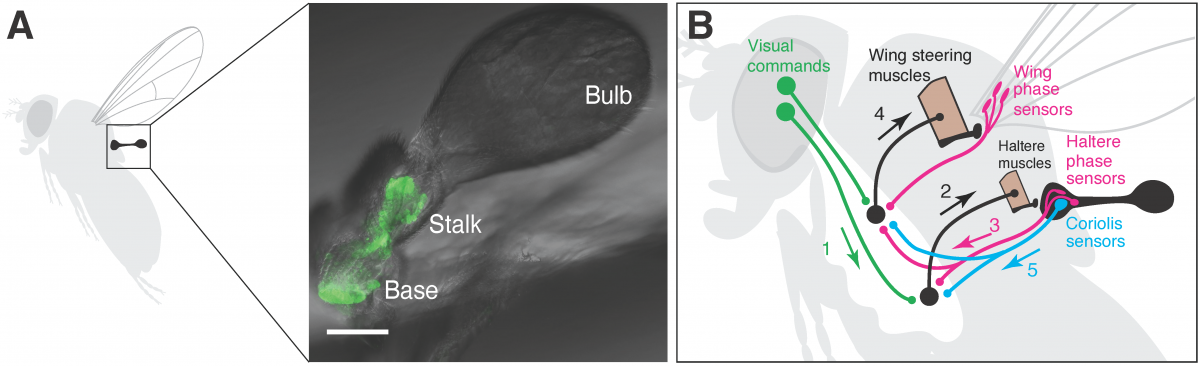
Figure: A) Location and structure of the Drosophila haltere. B) Schematic of the control loop linking visual and mechanosensory feedback for fly flight.
___________________________________________________________________________________________________________
Ellie Heckscher, Matthieu Louis, and Jane Loveless: Evolution of the neuromechanics of locomotion in the Drosophila larva
At the larval stage, Drosophila move through waves of peristaltic contractions that travel from the posterior (tail) to the anterior (head) end of the body. The execution of these waves is rhythmically organized across body segments. While symmetrical contractions produce forward or backward locomotion, asymmetric contractions underlie turning maneuvers and active sampling through sweeping movements of the head. How peristalsis emerges from the interplay between the biomechanics of the larval musculature and its nervous system remains elusive. We will approach this question by comparing the locomotor behaviors and neuromusculatures of different species of the Drosophila group (and beyond). In particular, we will examine whether and how evolution has modified the control of peristaltic locomotion to adapt to the environmental properties of distinct ecological niches.
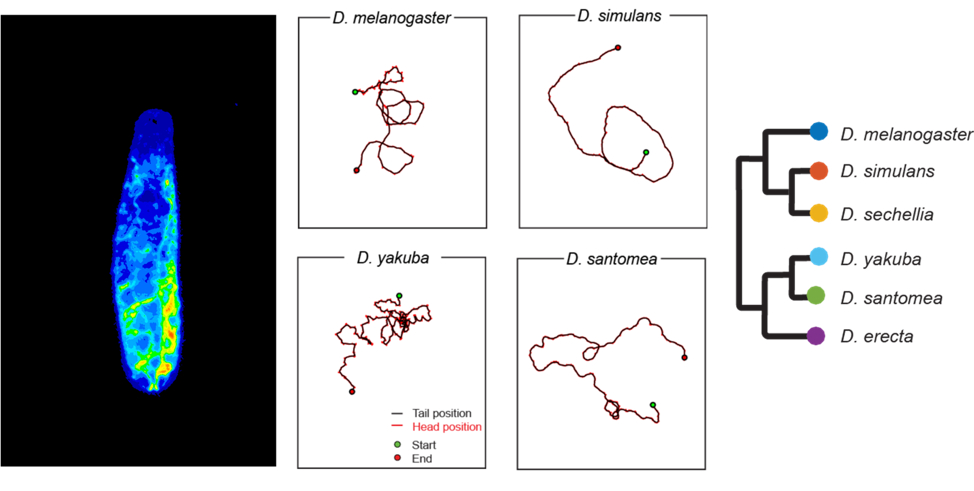
Figure: Drosophila larva (left) and examples of larval trajectories of four Drosophila species on an agarose substrate with the same stiffness (Elie Fink, Louis lab).
___________________________________________________________________________________________________________
Jimmy Liao and Claire Wyart: Long-scale patterns in the kinematics of fish during navigation, hydrodynamics of schooling, and optomotor responses
Larvae and juvenile zebrafish exhibit robust orientation behaviors in response to visual gradients and water flow. In this module, we will investigate the long-scale kinematics of locomotion of fish navigating in visual or flow gradients to understand how patterns emerge in either single individuals or schools of fish. We will compare the effects of the presence of conspecifics to visual and acoustovestibular stimuli eliciting escape responses in single individuals. Building on this analysis, we will examine how the presence of other conspecifics influence the long-scale dynamics displayed by individuals.
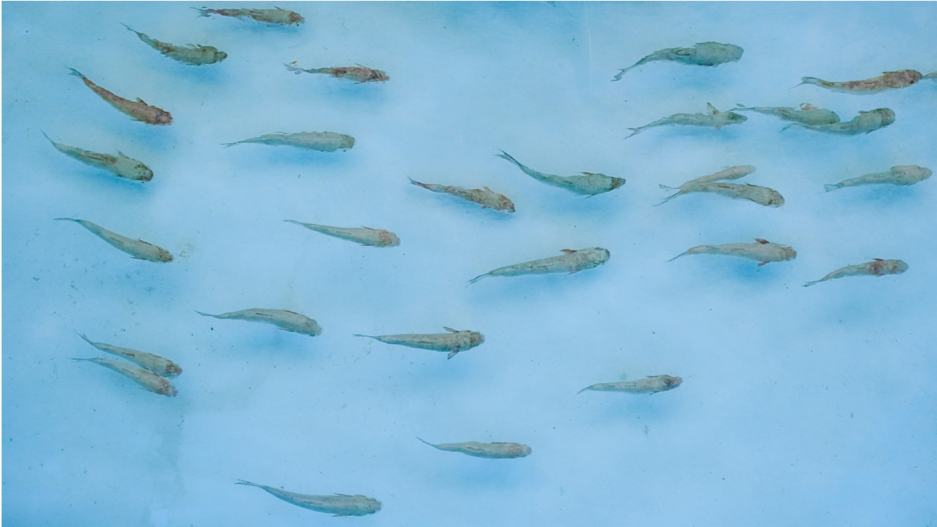

![]()
___________________________________________________________________________________________________________
Eugenia Chiappe, Pavan Ramdya, Julie Simpson, and Andrew Straw: How Flies Walk: exploring behavioral algorithms for sensory control of legged locomotion over complex terrain using adult Drosophila
Flies coordinate six legs to walk, cross gaps, and climb over obstacles. They display a range of gaits and exploratory movements but the full behavioral repertoire and underlying neural circuits are not fully understood. Visual, mechanosensory, and proprioceptive inputs contribute to the limb coordination required. We will assess flies’ capacity for motor learning using an obstacle course or treadmill to measure spatial memory and locomotor strategies under different sensory conditions. This module will use quantitative behavioral analyses and optogenetic perturbations to probe the mechanisms of walking in flies, in particular how sensory feedback and central pattern generators may underlie locomotor control.
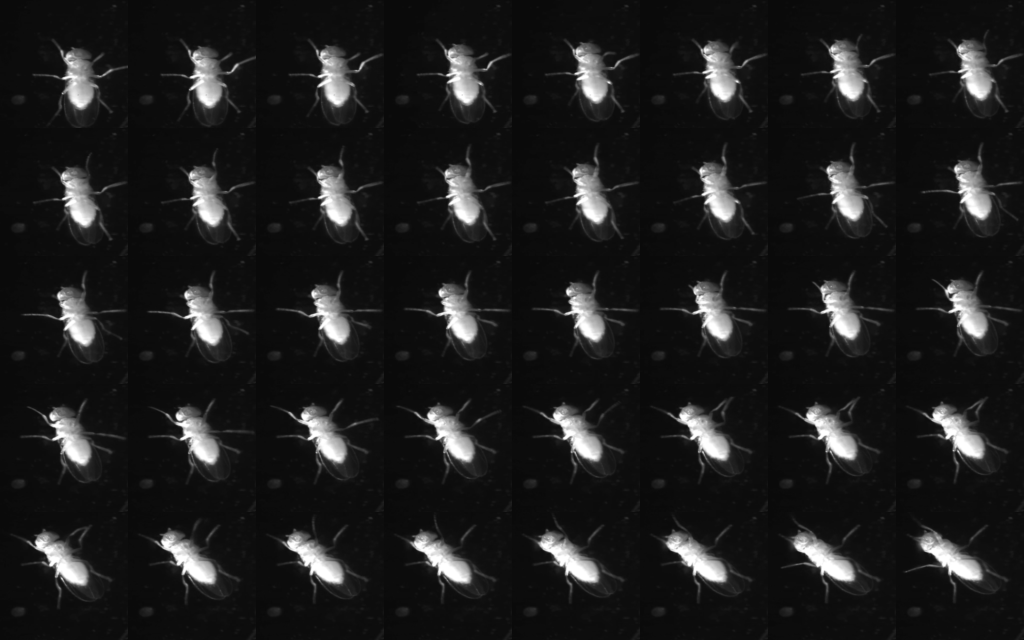
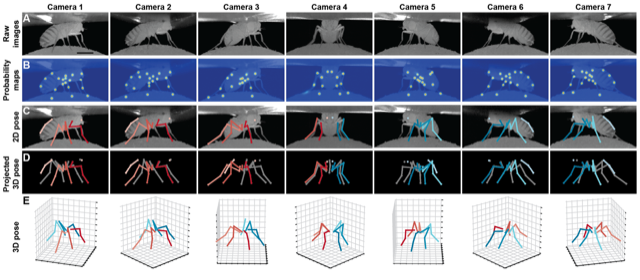

Walking fly figures: Ramdya lab (top) and Chiappe lab (bottom)
